Experiments in Agentic AI, Part V: Building a Knowledge Pipeline
by Alex Harvey
- Introduction
- Architecture
- The Code
- The Ingestion Worker
- The Indexing Job
- The Retrieval API
- Running the Pipeline
- What Comes Next
Introduction
Today’s post in my series documenting my experiments in Agentic AI is an exciting departure from the previous parts (I — IV).
Until now, I had been experimenting with multi-agent AI workflows, on the assumption that the interesting engineering problem lay in getting AI agents to collaborate effectively. The system I built included planners, researchers, analysts, writers, and reviewers collaborating in the production of a research report on Beatles song lyrics.
But over time I came to understand that the harder problem lies elsewhere: the retrieval system that feeds data to the agents.
In the earlier posts I used LangChain (Parts I–III) and then, in Part IV, I swapped in LlamaIndex, which proved easier to work with and helped clarify how indexing and retrieval should be structured.
However, those experiments still ran mostly as local pipelines designed for exploration rather than deployment.
Today I present an entirely new stack — one designed to be deployable in a real-world environment. The system runs as an AWS pipeline, with containerised services in ECS, persistent storage in S3 and RDS, and deployment orchestrated through GitHub Actions.
So, this post is not about clever agent design, but about building the infrastructure that makes reliable AI workflows possible.
Architecture
The overall architecture of the system is shown below.
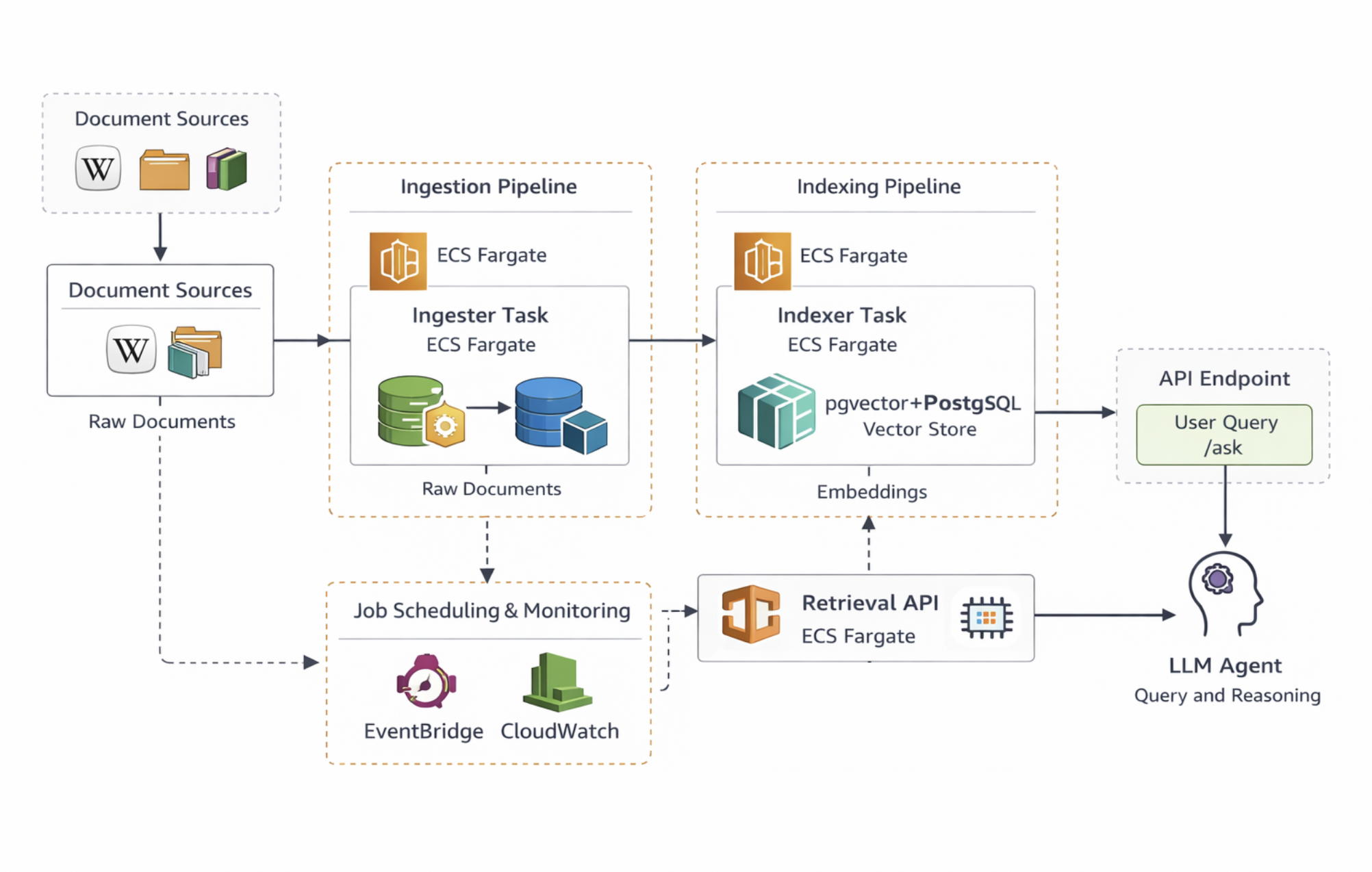
At a high level, the architecture has three layers:
- Ingestion pipeline
- Indexing pipeline
- Retrieval API + AI agent
Across all layers, I use ECS Fargate to run the API as well as the ingestion and indexing jobs inside Docker containers, avoiding the need to manage EC2 instances directly and the complexity of EKS and Kubernetes. The services themselves are packaged using Docker, which makes it easy to run the same containers locally during development and then deploy them unchanged in AWS.
The ingestion pipeline collects documents from Wikipedia, which I chose as a convenient stand-in for the kind of internal knowledge bases many organisations maintain. In most companies, AI systems are expected to answer questions based on internal documentation or a corporate wiki, and Wikipedia provides a useful proxy: it is large, structured as interlinked pages, and similar in style to internal knowledge repositories such as Confluence.
Raw documents collected during ingestion are stored in an S3 bucket, providing cheap and durable storage for the corpus while keeping the original data separate from the indexing process.
LlamaIndex is the only component retained from Part IV. It is used to parse documents, split them into chunks, and construct the retrieval structures used by the system. Each chunk of text is converted into a vector embedding using an OpenAI embedding model, allowing semantic similarity search to be performed later.
These embeddings are stored in an RDS PostgreSQL database. Using pgvector keeps the system architecture simple by allowing vector search to run inside PostgreSQL rather than introducing a separate vector database service.
The serving layer is implemented using FastAPI, which exposes a simple API endpoint that handles user questions and performs retrieval.
Behind the API sits a single LLM agent (OpenAI), whose role is to retrieve relevant context and generate an answer grounded in the retrieved documents.
The ingestion and indexing jobs are scheduled using EventBridge, allowing the system to periodically refresh its data without manual intervention.
Finally, GitHub Actions handles the build and deployment pipeline, while CloudWatch collects logs and metrics so the behaviour of the system can be monitored.
The Code
The full code for the system described in this post is available on GitHub:
https://github.com/alex-harvey-z3q/wiki-rag
The repository contains the infrastructure, ingestion workers, indexing pipeline, and API service used to run the system.
The simplified structure of the project looks as follows:
api/ FastAPI retrieval service
indexer/ LlamaIndex indexing pipeline
ingest/ Wikipedia ingestion worker
terraform/ AWS infrastructure
scripts/ Deployment and utility scripts
The Ingestion Worker
The ingestion worker is responsible for collecting documents and storing them in S3 so they can be indexed later.
For this project, the ingestion step retrieves articles from Wikipedia. The goal is not to index the entirety of Wikipedia, but rather to build a representative corpus that behaves similarly to the kind of internal documentation systems commonly found in organisations.
The ingestion worker runs as a containerised ECS task and performs a relatively simple workflow:
- retrieve documents from Wikipedia
- convert them into a structured format
- write them to S3
The worker deliberately performs minimal processing. Its purpose is simply to collect and persist the raw data. More complex processing is deferred to the indexing stage.
A simplified version of the ingestion process looks like this:
documents = loader.load_data()
for doc in documents:
s3.put_object(
Bucket=bucket,
Key=f"docs/{doc.id}.json",
Body=json.dumps(doc)
)
By storing the original documents in S3, the system preserves the raw corpus independently of the indexing pipeline. If the indexing logic changes in the future — for example, if a different chunking strategy or embedding model is used — the index can simply be rebuilt from the stored documents.
The Indexing Job
Once documents have been collected and stored in S3, the indexing job processes them to build the vector search index.
This job also runs as an ECS task and uses LlamaIndex to handle document parsing, chunking, and embedding. During indexing, the documents are split into smaller chunks of text, which are then converted into vector embeddings.
These embeddings are generated using an OpenAI embedding model and stored in PostgreSQL using the pgvector extension.
In LlamaIndex, the indexing step can be expressed quite succinctly:
index = VectorStoreIndex.from_documents(
documents,
storage_context=storage_context
)
Behind the scenes, this process performs several important steps:
- splitting documents into chunks
- generating embeddings for each chunk
- storing the embeddings and metadata in the database
The result is a vector index that can be queried using semantic similarity search.
Separating the indexing stage from ingestion allows the pipeline to scale more effectively. Documents can be collected independently of indexing, and indexing can be rerun whenever necessary without re-fetching the source data.
The Retrieval API
The final layer of the system is the retrieval API, which serves user queries.
This layer is implemented using FastAPI and runs as a long-lived ECS service. The API exposes a simple endpoint that accepts a question and returns an answer generated by the language model.
A simplified version of the endpoint looks like this:
@app.post("/ask")
async def ask(question: str):
response = query_engine.query(question)
return {"answer": str(response)}
When a request arrives, the system performs several steps:
- the user’s question is converted into an embedding
- the embedding is used to perform a vector similarity search
- the most relevant chunks of text are retrieved from the database
- these chunks are assembled into a prompt
- the language model generates a final response
The retrieved context grounds the model’s response in the indexed documents, allowing it to answer questions based on the underlying corpus rather than relying solely on its pre-trained knowledge.
In contrast to the earlier parts of this series, the agent used here is intentionally simple. Instead of coordinating multiple specialised agents, the system relies on a single language model that retrieves context and produces an answer.
This simplification is deliberate. The goal of this experiment is not to build the most complex agent workflow possible, but rather to build the infrastructure required to support complex Agentic AI workflows.
Running the Pipeline
With the ingestion worker, indexing job, and retrieval API in place, the system forms a simple but complete knowledge pipeline.
The pipeline runs in three stages.
First, the ingestion job retrieves documents from Wikipedia and stores them in S3. This stage is responsible only for collecting and preserving the raw data.
Second, the indexing job reads the stored documents, splits them into chunks, generates embeddings using the OpenAI embedding model, and writes the resulting vectors into the PostgreSQL database using pgvector.
Finally, the FastAPI service exposes a query endpoint that retrieves relevant document chunks and generates responses using the language model.
Because the ingestion and indexing stages run as independent ECS tasks, they can be triggered periodically using EventBridge. This allows the system to refresh its data automatically without interrupting the retrieval API.
In practice, the operational flow looks like this:
- ingestion task runs and collects documents
- documents are written to S3
- indexing task processes the documents
- embeddings are stored in PostgreSQL
- the API serves queries using the indexed data
This separation makes the system easier to reason about. Each stage performs a single task, and failures in one part of the pipeline do not necessarily bring down the rest of the system.
What Comes Next
This experiment focused primarily on building the infrastructure required to support a retrieval-augmented AI system.
While the agent itself is intentionally simple, the pipeline now provides a foundation that could support more sophisticated workflows in the future.
In Part VI, I plan to explore how more sophisticated agents might interact with a system like this, and how retrieval quality can be evaluated and improved over time.
For now, however, the key takeaway from this experiment is that building useful AI agents often begins not with the agents themselves, but with the data pipelines that feed them.
tags: rag - agentic-ai